Overview
antedep supports three model families:
- AD (Gaussian) for continuous longitudinal outcomes
- INAD for count longitudinal outcomes
- CAT (categorical AD) for discrete state longitudinal outcomes
Packaged datasets include bolus_inad,
cattle_growth, cochlear_implant,
labor_force_cat, and race_100km.
Production-Readiness Matrix
| Model | Data type | Complete-data fit/logLik | Missing-data fit/logLik | Notes |
|---|---|---|---|---|
| AD | Continuous | Ready | Ready (fit_gau, logL_gau) |
Missing-data fit supports EM / observed-data likelihood modes |
| INAD | Counts | Ready | Ready (fit_inad, logL_inad) |
Missing-data fit supports
na_action = "marginalize"
|
| CAT | Categorical states | Ready | Ready (fit_cat, logL_cat) |
Missing-data fit supports orders 0, 1, 2 |
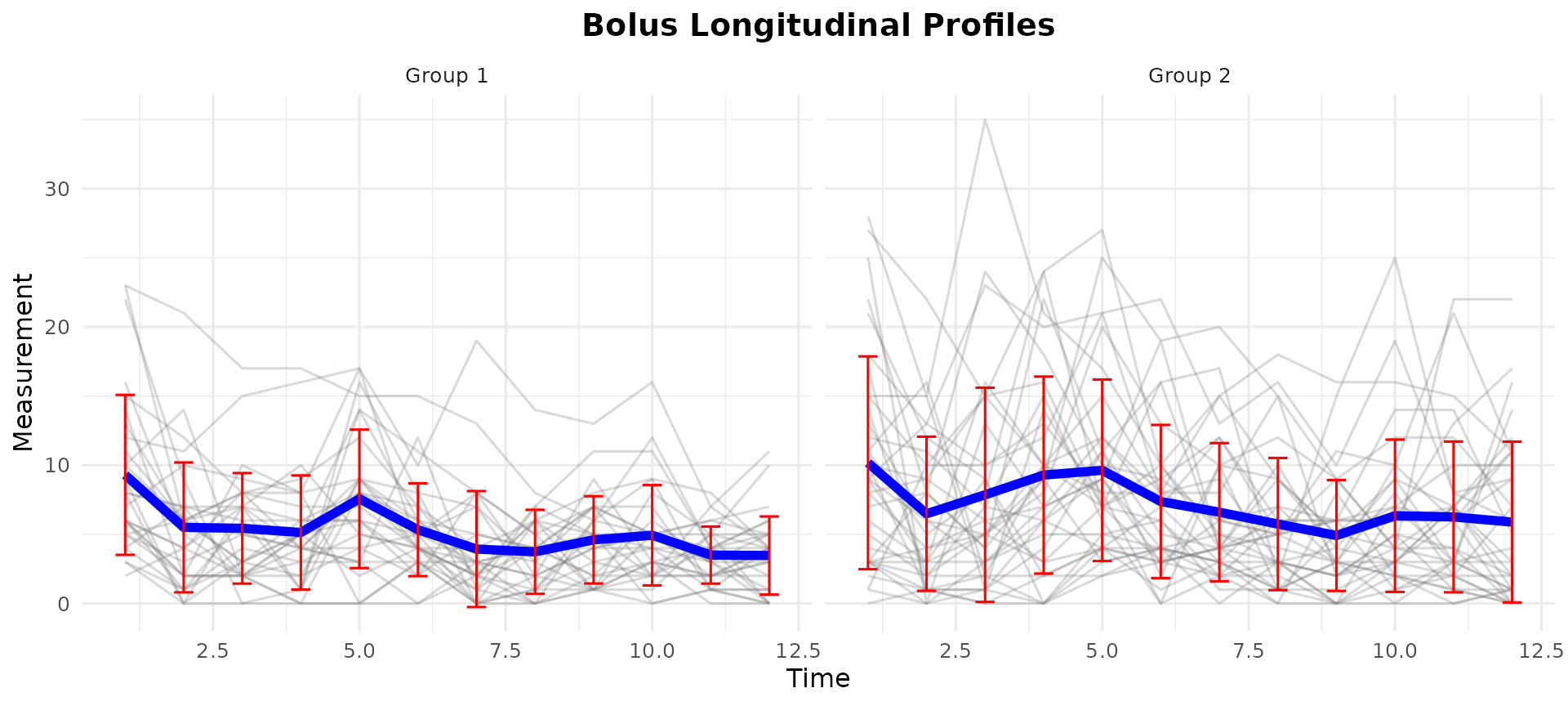
Visualizing Longitudinal Data
data("bolus_inad")
plot_profile(
bolus_inad$y,
blocks = bolus_inad$blocks,
block_labels = c("Group 1", "Group 2"),
title = "Bolus Longitudinal Profiles"
)
# More detailed dependence diagnostic
plot_prism(bolus_inad$y)AD Workflow (Continuous Outcomes)
Complete-data fit and order comparison
y_gau <- simulate_gau(n_subjects = 20, n_time = 4, order = 1, phi = 0.5)
fit_gau1 <- fit_gau(y_gau, order = 1)
fit_gau2 <- fit_gau(y_gau, order = 2)
c(order1_logLik = fit_gau1$log_l, order2_logLik = fit_gau2$log_l)
#> order1_logLik order2_logLik
#> -105.6720 -104.4205
bic_order_gau(y_gau, max_order = 2)$table
#> order log_l bic selected
#> order0 0 -112.6549 249.2756 FALSE
#> order1 1 -105.6720 244.2971 TRUE
#> order2 2 -104.4205 247.7855 FALSE
# Example AD order test
test_order_gau(y_gau, p = 0, q = 1)$p_value
#> [1] 0.002952214Missing-data AD fit
y_gau_miss <- y_gau
y_gau_miss[sample(length(y_gau_miss), 8)] <- NA
fit_gau_em <- fit_gau(y_gau_miss, order = 1, na_action = "em", em_max_iter = 10)
fit_gau_cc <- fit_gau(y_gau_miss, order = 1, na_action = "complete")
#> Warning: Only 13/20 subjects have complete data (65%).
#> Consider using na_action = 'em' for less information loss.
c(em_logLik = fit_gau_em$log_l, complete_case_logLik = fit_gau_cc$log_l)
#> em_logLik complete_case_logLik
#> -94.00107 -63.11882
fit_gau_em$pct_missing
#> [1] 10AD mean and covariance structure tests (complete data)
These AD inference tests currently require complete data.
# One-sample and two-sample mean-profile tests
mu0 <- colMeans(y_gau)
blocks_gau <- rep(1:2, each = nrow(y_gau) / 2)
test_one_sample_gau(y_gau, mu0 = mu0, p = 1)$p_value
#> [1] 1
test_two_sample_gau(y_gau, blocks = blocks_gau, p = 1)$p_value
#> [1] 0.01854549
# Wald contrast test: H0 that adjacent means are equal
C <- diff(diag(ncol(y_gau)))
test_contrast_gau(y_gau, C = C, p = 1)$p_value
#> [1] 0.8183341
# Covariance homogeneity across groups
test_homogeneity_gau(y_gau, blocks = blocks_gau, p = 1)$p_value
#> g1
#> 0.3237498INAD Workflow (Count Outcomes)
Complete-data INAD fit
y_inad <- simulate_inad(
n_subjects = 20,
n_time = 4,
order = 1,
thinning = "binom",
innovation = "pois",
alpha = 0.4,
theta = 3
)
fit_inad1 <- fit_inad(y_inad, order = 1, thinning = "binom", innovation = "pois")
fit_inad1$log_l
#> [1] -158.3541
bic_order_inad(y_inad, max_order = 2, thinning = "binom", innovation = "pois")$table
#> order logLik n_params BIC
#> order_0 0 -164.6683 4 341.3196
#> order_1 1 -158.3541 7 337.6784
#> order_2 2 -158.3541 8 340.6741Missing-data INAD fit
y_inad_miss <- y_inad
y_inad_miss[sample(length(y_inad_miss), 10)] <- NA
fit_inad_miss <- fit_inad(
y_inad_miss,
order = 1,
thinning = "binom",
innovation = "pois",
na_action = "marginalize",
max_iter = 10
)
fit_inad_miss$log_l
#> [1] -142.2786
fit_inad_miss$pct_missing
#> [1] 12.5
# Missing-data LRTs (observed-data likelihood)
blocks_inad <- rep(1:2, each = nrow(y_inad_miss) / 2)
test_order_inad(y_inad_miss, order_null = 0, order_alt = 1, blocks = blocks_inad)$p_value
#> [1] 0.08052965
test_homogeneity_inad(y_inad_miss, blocks = blocks_inad, order = 1, test = "mean")$p_value
#> [1] 0.7852657CAT Workflow (Categorical Outcomes)
Complete-data CAT fit and inference
y_cat <- simulate_cat(n_subjects = 20, n_time = 4, order = 1, n_categories = 3)
fit_cat1 <- fit_cat(y_cat, order = 1)
fit_cat1$log_l
#> [1] -79.08119
# Wald CIs (complete data)
ci_cat1 <- ci_cat(fit_cat1, parameters = "marginal")
head(ci_cat1$marginal, 3)
#> parameter time category estimate se lower upper level
#> cat_1 pi_t1_cat1 1 1 0.25 0.09682458 0.0602273 0.4397727 0.95
#> cat_2 pi_t1_cat2 1 2 0.40 0.10954451 0.1852967 0.6147033 0.95
#> cat_3 pi_t1_cat3 1 3 0.35 0.10665365 0.1409627 0.5590373 0.95
# Order tests for CAT
run_order_tests_cat(y_cat, max_order = 2)$table
#> comparison order_null order_alt method lrt_stat df p_value significant
#> 1 AD(0) vs AD(1) 0 1 lrt 13.04437 12 0.36582405 FALSE
#> 2 AD(1) vs AD(2) 1 2 lrt 37.28660 24 0.04096144 TRUEMissing-data CAT fit
y_cat_miss <- y_cat
y_cat_miss[sample(length(y_cat_miss), 12)] <- NA
fit_cat_miss <- fit_cat(y_cat_miss, order = 1, na_action = "marginalize")
fit_cat_miss$log_l
#> [1] -65.91133
fit_cat_miss$settings$na_action_effective
#> [1] "marginalize"
# EM alternative for CAT missing data (orders 0/1)
fit_cat_miss_em <- fit_cat(y_cat_miss, order = 1, na_action = "em", em_max_iter = 10)
#> Warning in em_cat(y = y, order = p, blocks = blocks_input, homogeneous =
#> homogeneous, : EM did not converge after 10 iterations (change = 0.000809).
fit_cat_miss_em$log_l
#> [1] -65.91256
# Missing-data order/homogeneity tests are supported
blocks_cat <- rep(1:2, each = nrow(y_cat_miss) / 2)
test_order_cat(y_cat_miss, order_null = 0, order_alt = 1)$p_value
#> [1] 0.2500927
test_homogeneity_cat(y_cat_miss, blocks = blocks_cat, order = 1)$p_value
#> [1] 0.3780152Additional Inference Helpers (Complete Data)
# INAD homogeneity/stationarity tools
blocks <- rep(1:2, each = nrow(y_inad) / 2)
run_homogeneity_tests_inad(y_inad, blocks = blocks, order = 1)
run_stationarity_tests_inad(y_inad, order = 1, blocks = blocks)
# CAT stationarity/time-invariance tools
test_timeinvariance_cat(y_cat, order = 1)
test_stationarity_cat(y_cat, order = 1)Known Limitations
- EM entry points:
em_gau(Gaussian),em_inad(INAD), andem_cat(CAT, orders 0/1) are available; for CAT order 2 with missing data, usefit_cat(na_action = "marginalize"). - For CAT missingness,
fit_cat(na_action = "marginalize")supports orders 0/1/2, whilefit_cat(na_action = "em")(orem_cat) supports orders 0/1 with optional safeguard step-halving. - Missing-data confidence intervals are not implemented yet
(
ci_gau,ci_inad,ci_catrequire complete-data fits). - AD missing-data LRT/mean/covariance tests are not implemented yet.
- CAT missing-data time-invariance/stationarity tests
(
test_timeinvariance_cat,test_stationarity_cat,run_stationarity_tests_cat) are not implemented yet. - Current order support is up to order 2 for AD, INAD, and CAT.
- CAT missing-data marginalization currently supports orders 0, 1, and 2.